One common frustration that I have heard expressed about R is that there is no automatic way to plot error bars (whiskers really) on bar plots. I just encountered this issue revising a paper for submission and figured I'd share my code. The following simple function will plot reasonable error bars on a bar plot.
[r]
error.bar <- function(x, y, upper, lower=upper, length=0.1,...){
if(length(x) != length(y) | length(y) !=length(lower) | length(lower) != length(upper))
stop("vectors must be same length")
arrows(x,y+upper, x, y-lower, angle=90, code=3, length=length, ...)
}
[/r]
Now let's use it. First, I'll create 5 means drawn from a Gaussian random variable with unit mean and variance. I want to point out another mild annoyance with the way that R handles bar plots, and how to fix it. By default, barplot() suppresses the X-axis. Not sure why. If you want the axis to show up with the same line style as the Y-axis, include the argument axis.lty=1, as below. By creating an object to hold your bar plot, you capture the midpoints of the bars along the abscissa that can later be used to plot the error bars.
[r]
y <- rnorm(500, mean=1)
y <- matrix(y,100,5)
y.means <- apply(y,2,mean)
y.sd <- apply(y,2,sd)
barx <- barplot(y.means, names.arg=1:5,ylim=c(0,1.5), col="blue", axis.lty=1, xlab="Replicates", ylab="Value (arbitrary units)")
error.bar(barx,y.means, 1.96*y.sd/10)
[/r]

Now let's say we want to create the very common plot in reporting the results of scientific experiments: adjacent bars representing the treatment and the control with 95% confidence intervals on the estimates of the means. The trick here is to create a 2 x n matrix of your bar values, where each row holds the values to be compared (e.g., treatment vs. control, male vs. female, etc.). Let's look at our same Gaussian means but now compare them to a Gaussian r.v. with mean 1.1 and unit variance.
[r]
y1 <- rnorm(500, mean=1.1)
y1 <- matrix(y1,100,5)
y1.means <- apply(y1,2,mean)
y1.sd <- apply(y1,2,sd)
yy <- matrix(c(y.means,y1.means),2,5,byrow=TRUE)
ee <- matrix(c(y.sd,y1.sd),2,5,byrow=TRUE)*1.96/10
barx <- barplot(yy, beside=TRUE,col=c("blue","magenta"), ylim=c(0,1.5), names.arg=1:5, axis.lty=1, xlab="Replicates", ylab="Value (arbitrary units)")
error.bar(barx,yy,ee)
[/r]

Clearly, a sample size of 100 is too small to show that the means are significantly different. The effect size is very small for the variability in these r.v.'s. Try 10000.
[r]
y <- rnorm(50000, mean=1)
y <- matrix(y,10000,5)
y.means <- apply(y,2,mean)
y.sd <- apply(y,2,sd)
y1 <- rnorm(50000, mean=1.1)
y1 <- matrix(y1,10000,5)
y1.means <- apply(y1,2,mean)
y1.sd <- apply(y1,2,sd)
yy <- matrix(c(y.means,y1.means),2,5,byrow=TRUE)
ee <- matrix(c(y.sd,y1.sd),2,5,byrow=TRUE)*1.96/sqrt(10000)
barx <- barplot(yy, beside=TRUE,col=c("blue","magenta"), ylim=c(0,1.5), names.arg=1:5, axis.lty=1, xlab="Replicates", ylab="Value (arbitrary units)")
error.bar(barx,yy,ee)
[/r]

That works. Maybe I'll show some code for doing power calculations next time...
Clearly, I need to do something about these annoying line numbers in the text boxes and, for that matter, the text boxes themselves! Will work on that...
Thank you! Going right into my scripts collection.
Why do you divide 1.96*Standard Deviation by ten for your error bar?
Wouldn't 1.96* SD be the 95% confidence interval?
standard error = standard deviation/sqrt(N). The simulated data had N=100, therefore the 95% CI was approximately 1.96*sd/10.
That was really helpful, thank you!
I'm having a problem with the ylim, though. If I set it to be
ylim = c(450, 950)
then I get the axis correctly, but I still get the bars appearing as if they were starting at zero.
I don't know whether this is a limitation of ylim in barplot in itself or whether I'm doing something wrong.
Hard to know without the specifics of the problem, but it sounds like you didn't give the means as the y argument of the function.
How can i get the same CI errorbars when I have different size sample??
The bit of code that calculates the CIs is: ee <- matrix(c(y.sd,y1.sd),2,5,byrow=TRUE)*1.96/10. All you are doing is making a 2x5 matrix of the standard errors here by first constructing a matrix of standard deviations and then multiplying by z/sqrt(n). One very simple (if inelegant) way to plot error bars for two samples of different size it is to move the z/sqrt(n) inside the matrix() argument. Say that the size of one sample was 100 and the other was 144, then you could simply do this: ee <- matrix(c(y.sd*1.96/10,y1.sd*1.96/12),2,5,byrow=TRUE).
I have tried this but i get this message all the time
"Error: object 'error.bar' not found"
You need to source the code that I provide -- the function called "error.bar". Either copy the function (minus the annoying line numbers that the blog software adds) and paste it at the command line, save it to a file (e.g., error.bar.R) and source it, i.e., source("error.bar.R") (which assumes you saved the file in your working directory; otherwise, include the path), or use the capabilities of an R-aware text editor (e.g., ess mode in Emacs or Tinn-R). Once you have sourced it, the code should work.
how can one plot error bars when using "plot' with type='b'.
@sp: I don't know, but it would be similar to what I did with barplot() -- shouldn't be too hard.
Hi,
Thanks for demonstrating this; you are right, it is a major problem with R, particularly for new users like myself.
As this is going straight into a piece of work that I am currently doing, how do you prefered to be referenced?
Not really a citation. Put me in the acknowledgements.
Hello!
I tried your scrip but got an error message:
Error in error.bar(barx, yy, ee) : vectors must be same length
Thanks,
Ron
Well, you're three vectors do need to be the same length! barx, yy, and ee should all be of length k...
Hello,
Thanks a lot for your function.
I just have a problem though : I have already SE and means in my data, and when entering these arguments in the error.bar function
e.g. error.bar(graph, y_value, y_SE)
it says "unexpected input in "error.bar(barplot(etc..))"".
Do you have any idea why ?
Many thanks in advance,
Cheers
It's hard to know without seeing the exact inputs/outputs. I usually get the "unexpected input" error when I forget a comma between arguments, so I would check on that. At a minimum, you need three arguments, all separated by commas: error.bar(x,y,upper) [where 'upper' is the bit you want to add/subtract from your vector of means means, y]. This assumes symmetric CIs (which you can change by adding 'lower' as a separate argument).
Thank you very much, it works !
Now another challenge : I want to use the "sort" function to show the bars from the lowest to the highest y value. When doing it, the SD error bars are not sorted and don't match their y values.
How can I implement the sort function in the error .bar function ?
Many thanks in advance for your help !
Wow. I even got this to work with a line plot I was doing. It's a little artificial. instead of handing off the object assigned the line draw (like with what you did with barx <-) you can just hand the function a c() vector with the right number of points. Works like a charm. Thanks!
Thanks for this. Was looking for a way to plot barplots with errors before and totally happy to have found your explanation.
Hello, thank you so much for this information it's indeed very helpfull! I have only one question: how d0 you obtain "z" for the CI?
Cheers, Anna
Anna, not quite sure what you mean by "z" but I'll take a stab. I assume what you're asking is where did the 1.96 come from. This comes directly from the normal approximation.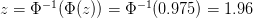 , where
, where 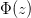 cumulative distribution function for a normal distribution, and
cumulative distribution function for a normal distribution, and 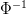 is its inverse. You know, it's that whole 95% of the observations in a normal distribution lie
is its inverse. You know, it's that whole 95% of the observations in a normal distribution lie 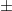 2 standard deviations from the mean? It's actually more like 1.96...
2 standard deviations from the mean? It's actually more like 1.96...
Great! Very useful!
Thanks!
Many thanks for the code! very useful!!
Great code! Very easy to use.
Thanks for posting this!
Thanks for this nice little code!
It works great; I also added the option to specify sd, se or CI for the error bars.
One thing I'm having trouble with is specifying ylim with a minimum other than zero. Any ideas?
Cheers,
Oops, posted the question too quickly. 'xpd=FALSE' fixed my problem with the bars extending below the axis.
Cheers!
Thank you so much. This was invaluable to me.
Just finished writing a postscript library to produce bar plots with error bars when I came across your R solution. Thanks for posting this! It's surprising how non-trivial it seems to find tools to do this...
thanks dear!!!
Thank you so much for posting this code, I lost so much time trying to get those error bars!
Yes, this is great! I have used it several times in reports. Thanks again. But do I need to cite you? And how?
plotCI() in the plotrix package
Thanks for writing this, way better than the way I was doing it previously with lines()!
Just a couple recommendations:
1) I added a lwd argument to the function so users can specify the line width.
2) I changed the standard error calculations to sqrt(length(COLUMN)) to get around the different sample size problem.
Best,
Chris
no need to cite, thanks.
Thank you very much for sharing the code! Unfortunatelly, I couldnt make the code work for a boxplot with "beside=F". Do you have any idea?Best,
Sabine
Thanks for your help!
I would like to modifie the function in order to have a smaller arrow. how do I do it?
ciao
Matteo
Sorry, not much to go on here. I hope it worked out for you...
Look at the help file for arrows(). There are "length", "width", "angle", and "code" arguments that allow you to change the style of the arrow head.
Great function--this helped me out greatly! Thanks so much for posting. It is often these short functions that can save tons of time...
Thank you
Thanks!! Will use this all the time.
Just wanted to add my thanks for posting this. It's exactly what I was looking for
Very elegant way to do it! As shown in the examples, it works well for bar plots. For point plots I want to rub out the middle of the arrow so that it doesn't overlap my plotting symbol. To do this I plotted the error bars first and then plotted my points with pch = 21 and bg = "white". Does anyone have a more elegant or more general method?
That's probably what I would do!
I used this coed to produce error bars in a "side by side" bar plot (like the second graph in your example). While the code works and produced error bars, they are mysteriously divorced from the values in the bars; some of them float above the bars and others are completely embedded somewhere within the bar. Any thoughts?
Hard to know without more specifics. My guess is that your vectors don't match up so that you're getting weird effects of recycling. This is a natural way that error bars might get offset from the means.
Hi, great post, thanks heaps! I'm wondering how I need to adapt the code to help me calculate the appropriate confidence intervals when I have a poisson response variable and factor predictor variable...If you have some quick hints, they are greatly appreciated!
Awesome post.